We are a multidisciplinary international team of nine academic and industrial partners.
By our cooperation we fill major technical and scientific gaps to allow a full-fledged application of contemporary synthetic biology to large-scale industrial biotechnology in Europe.
1. Wageningen University
 Wageningen University
Wageningen University
Centre for Systems & Synthetic Biology (WCSB)
Wageningen, The Netherlands
The Wageningen University (WU) is an internationally leading education and research organisation. As a part of its strategic plan, the WU has set Systems & Synthetic Biology as one of its spearheads, having established in 2010 the Chair of Systems and Synthetic Biology and, in 2012, the overarching Wageningen Centre for Systems & Synthetic Biology (WCSB). The department of Systems and Synthetic Biology contributes to the elucidation of the mechanisms underlying basic cellular processes, evolution and interactions among microbes and between microbes and their environment and to translate this knowledge into applications of biotechnological, medical and environmental interest. WCSB, set up to coordinate all systems biology activities at Wageningen University, will be the primary cooperation partner for research centres and industry in this specialist field. The WCSB’s research focuses on the entire spectrum of biological systems, from DNA to ecosystem, and involves topics relevant to plants, digestion and microbes. Both are headed by Prof. Martins dos Santos.
 Prof. Dr. Dipl.-Ing. Vítor A. P. Martins dos Santos
Prof. Dr. Dipl.-Ing. Vítor A. P. Martins dos Santos
Main Contact
Prof. Dr. Dipl.-Ing. Vítor Martins dos Santos, Chair of Systems and Synthetic Biology and Head of the Wageningen Centre for Systems & Synthetic Biology is also the president of the Dutch Society of Biotechnology (NBV). His lab develops and applies theoretical frameworks supporting experimental research towards the systems level understanding of the hierarchies and dynamics of cellular networks. These frameworks are used for directed cellular reprogramming, with applications to health and industry. The lab has made acknowledged contributions to large-scale genome engineering of designer microbes and to cellular-reprogramming.
| Modellers: | Experimentalists: |
|
Nhung Pham MSc Ruben van Heck MSc |
Linde Kampers MSc Stamatios Damalas MSc |
The missing link between computer and laboratory
Model-driven metabolic engineering of Pseudomonas putida
Microscopic animals called microorganisms are used to produce diverse products, including, but not limited to, food additives, antibiotics, vaccines, bioplastics, bio-pesticides and biofuels. Often, the use of microorganisms enables a much greener, faster and cheaper alternative to produce products in high volumes than conventional chemical processes. Each species of microorganism used has its own benefits, but also downsides. With the techniques we now have available, it is possible to choose the ideal candidate, and then improve it so that it has little to no downsides.
Within our EmPowerPutida project, we have chosen for Pseudomonas putida as the best candidate. It can withstand much stress, which will allow it to flourish and produce more than other bacteria on a large scale. It is also able to adapt quickly to new surroundings. To top it off, Pseudomonas putida is Generally Regarded as Safe (GRAS), which means it has passed extensive safety tests with flying colours.
Nowadays, this great method can be coupled to the world of computers. You can predict accurately how much effect an improvement can have. This allows for less wasting of time and money than if all ideas are directly tried in the lab. The results gathered at the lab can in turn be used to improve computer predictions. Often, within an organisation, there is only a limited amount of people proficient with making the computer predictions, opposed to many who experiment on the lab.
At Wageningen University & Research, the Systems and Synthetic Biology group, headed by Prof. Dr. Ir. Vitor Martins dos Santos, we go beyond just small-scale collaborations. Instead, we focus on building highly interactive bridges between dry computer predictions (dry lab) and wet laboratory experiments (wet lab). The number of dry lab workers is about equal to the number of wet lab workers. This allows for us to apply predictions where they haven’t been applied before, finding new solutions to yet unsolved problems. This close collaboration between people also enables the direct and fast exchange of new results gathered on the lab to feed back into the predictions, a process which can otherwise take months.
Within an eternal feedback loop, we use the information gained from metabolic models to optimize experimental setup, and use experimental data to improve the existing models. By combining these two techniques, we show that the sky is the limit. Within the EmPowerPutida project, we improve Pseudomonas putida in many ways to increase its potential as the workhorse for biocatalysis on an industrial level.
The state-of-the-art tools and methods in systems biology provide opportunities to understand and redesign the metabolism of Pseudomonas putida. In the EmPowerPutida Project, we use genome scale metabolic models as a platform to design metabolic engineering strategies for the production of native and non-native compounds in Pseudomonas putida. In silico (dry lab) tools are used to pinpoint possible targets for gene knock-out, knock-in or other genetic modifications that overcome bottlenecks in biochemical production processes. This results in a change of metabolic fluxes from growth to production. The in silico design is then implemented in the wet lab.
At the wet lab, implementation includes improvement and further development o
f a plug-and-play synthetic assembly system to edit the Pseudomonas putida genome, but also includes combining gene insertion with evolutionary pressure to increase its tolerance to grow under multiple different conditions. An example of this is increasing the bacterial survival rates under low oxygen conditions.
The question is not what we at Wageningen UR do to empower Pseudomonas putida. The question is how you would like to empower it!

The Wageningen EmPowerPutida team. From left to right: Linde Kampers MSc, Ruben van Heck MSc, Nhung Pham MSc, Stamatios Damalas MSc and Prof. Dr. Ir. Vitor Martins dos Santos.
2.1. Universitaet Stuttgart - Institute of Biochemical Engineering

Universitaet Stuttgart
Institute for Biochemical Engineering (IBVT)
Stuttgart, Germany
The Institute of Biochemical Engineering (Institut für Bioverfahrenstechnik, IBVT, University of Stuttgart) is focused on bioprocess development based on systems metabolic engineering and synthetic biology. Microbial and mammalian cells are studied thoroughly applying metabolomics and mRNA analysis for modelling and model-based strain development.
As such novel production platforms (e.g. for Escherichia coli) are created and further elaborated as new microbial producers. The interaction of global regulatory networks is elucidated while mirroring large-scale production conditions. Reasons and consequences of population heterogeneity – especially occurring under large-scale stress conditions – are unraveled.
 Prof. Dr.-Ing. Ralf Takors
Prof. Dr.-Ing. Ralf Takors
Main Contact
Prof. Ralf Takors is heading the IBVT since July 2009 after he had worked at Evonik Industries AG for about 5 years. There he was responsible for metabolic engineering and bioprocess development from lab- to production scale. In 2004 he had finished his 7-year lasting ‘Habilitation’ at Forschungszentrum Juelich/RWTH Aachen (Germany). Ralf is trained as a process engineer and has received Dr.-Ing. for Biochemical Engineering in 1997.
Dr. Bastian Blombach studied Biotechnology at the University of Applied Sciences Weihenstephan with internships at Creatogen AG and Boehringer Ingelheim (Vienna). During his Ph.D. he worked at the Institute of Microbiology and Biotechnology of Ulm University. Since 2012 he is junior group leader of the ‘Molecular Biotechnology’ group at the IBVT.
2.2. Universitaet Stuttgart - Institute of Technical Biochemistry
 Universitaet Stuttgart
Universitaet Stuttgart
Institute of Technical Biochemistry (ITB)
Stuttgart, Germany
The Institute of Technical Biochemistry at the Universitaet Stuttgart is led by Prof. Dr. Bernhard Hauer and covers biocatalysis, functional biomaterials, molecular biotechnology, fermentation and bioinformatics. The interdisciplinary research focuses on the cloning, expression, characterization and optimization of enzymes including P450 monooxygenases, lipases, enoate reductases, dehydratases and cyclases. Methods of laboratory evolution are used to generate novel and improved biocatalysts for industrial applications. The laboratories are equipped to implement analytical chemistry, microbiology, protein biochemistry and molecular biology. The research requires contributions from many disciplines, including chemistry, biochemistry, molecular biology, microbiology, bioengineering and bioinformatics.
 Prof. Dr. Bernhard Hauer
Prof. Dr. Bernhard Hauer
Main Contact
Prof. Dr. Bernhard Hauer has spent most of his career in industry working for BASF SE in Ludwigshafen, Germany, where he led the white biotechnology efforts. He has a strong background in this field and brings insights into industrial research and process development. He is the author of more than 200 articles and patents covering various topics of biotechnology and specialised areas. Since 2009 he is the head of the Institute of Technical Biochemistry at the Universitaet Stuttgart, Germany. He has been awarded with the BASF Award for Innovation of BASF SE (1997), the Dechema Award of the Max Buchner Research Foundation (2003) and the Biocat Award (2004).
The Institute of Technical Biochemistry is convinced that catalysis using microorganisms as well as isolated enzymes has a considerable influence on the synthesis of novel attractive products by developing greener manufacturing processes. This can only be achieved through a close collaboration between scientists from different fields. Our research requires contribution from many disciplines including chemistry, biochemistry and biology. We work in the spirit of discovery and enquiry in the functions and purposes of living catalysts namely enzymes and thus, are interested to understand the workings of all these enzymatic catalysts. The last years have witnessed key shifts in biotechnology aiming to present an attractive alternative to chemical catalysts. We have always anticipated these trends, pioneering research expanding the range of accessible reactions with novel enzymes and offer new synthesis routes to target compounds, some of which are given below.
Research focuses on the identification and development of novel enzymes allowing to further expand the toolbox of available biocatalysts to produce molecules of interest. Here, following enzyme-catalyzed reactions are being investigated:
- Selective oxyfunctionalizations
- Cytochrome P450 Monooxygenase
- Dioxygenases
- Enzymes for proton-initiated reactions
- Squalene-Hopene Cylcases
- Stereo- and regioselective water addition and elimination
- (De-)hydratases
- Asymmetric bioreductions of imines and activated alkenes
- Imine Reductases
- Ene Reductases
- Selective methylations and alkylation
- N-, O- and C-Methyltransferases
At the Institute of Technical Biochemistry, two PhD-students, Rebecca Demming (M. Sc.) and Max-Philipp Fischer (M. Sc.), are currently working on the enzymatic dehydration of short-chain alcohols and alkenols by the use of co-factor-independent bacterial (de‑)hydratases. Further, Dr. Wendy Escobedo is working as a Post-Doc on the indirect dehydration of short-chain alcohols via phosphorylation and dephosphorylation.
3. Consejo Superior de Investigaciones Científicas
 Consejo Superior de Investigaciones Científicas
Consejo Superior de Investigaciones Científicas
Madrid, Spain
The Spanish National Biotechnology Centre (Centro Nacional de Biotecnología, CNB) is ascribed to the Spanish National Research Council (Consejo Superior de Investigaciones Científicas, CSIC). At present, the CNB is the biggest research centre of the CSIC. The CNB stands out for its multidisciplinarity and encompasses two major scientific-technical areas within the CSIC. The centre’s departments of Macromolecular Structures, Cellular & Molecular Biology, Microbial Biotechnology and Immunology & Oncology belong to the area of Biology and Biomedicine; the department of Plant Molecular Genetics covers the area of Agricultural Sciences. In addition, a transversal Systems Biology Programme was initiated in 2009 with the vision to interconnect both areas.
 Prof. Dr. Víctor De Lorenzo
Prof. Dr. Víctor De Lorenzo
Main Contact
Professor of Research of the National Center of Biotechnology of the Spanish Research Council (CNB-CSIC) in Cantoblanco-Madrid (Spain). 325 Scientific publications at this time. Member of the Editorial Board of The Journal of Bacteriology (1989-2004). Member of the Editorial Board of Environmental Microbiology (1999-2008). Member of the Editorial Board of FEMS Microbiology Ecology (1998-2002). Member of the Editorial Board of Microbiology UK (mid 1999-2004). Member of the Editorial Board of Biodegradation (2001-2003). Member of the Editorial Board of Microbial Biotechnology (since 2008). Editor-in-Chief of Current Opinion in Biotechnology (since 2008). Member of the Editorial Board of Current Opinion in Microbiology (since 2008) Member of the Editorial Board of BioEssays (since 2009). Molecular Microbiology and Biotechnology. Genetic tools for Gram-negative bacteria destined for environmental release. Biodegradation of xenobiotic compounds. Environmental applications of recombinant antibodies, Synthetic Biology and Systems Biology. Metals in prokaryotic systems: Iron transport, resistances to heavy metals and metalloadsorption. Gene expression in Gram-negative bacteria, with focus in Pseudomonas. Regulation of catabolic pathways. Design of transcriptional factors and regulatory circuits á la carte.
Our laboratory is located at the Spanish National Center for Biotechnology, Madrid. In our laboratory, we are developing technologies to make biotechnology more applicable to environmental and industrial applications.
We are working with bacteria and developing technologies based on bacteria.
In order to adapt bacteria for use in industrial processes, the scientific community is focusing on several approaches. But first of all, how are bacteria used in industry? Bacteria can be used for biofuel production for our cars, for producing raw materials for medicines, and so on. Moreover, by using microorganisms we can do these processes cheaper and faster. In our laboratory, we focus on the model organism (the most preferred microorganism for a specific purpose), Pseudomonas putida (P. putida). P. putida's natural environment is soil. Each bacterial strain has its preferred living conditions as a result of evolution and it is called bacteria's natural environment. There are various bacteria types that have different natural environments. For example, P. putida is a soil bacterium, whereas Escherichia coli is a gut bacterium. How do we choose one as a model organism among all different bacterial strains then? It depends on the type of final applications one wants to realize. In our case for instance, we are benefiting from P. putida's evolutionary advantages of being a soil bacterium. Soil is a tough environment and requires adaptations for survival in harsh conditions. For this reason P. putida makes as a good starting point for industrial applications.
We are focusing our investigations on both the fundamental and application level. The knowledge we get from the fundamental level gives us a deeper understanding of the processes in bacteria as whole entities. This knowledge can then be used to provide robust and effective solutions in a variety of applications.
The applications in our laboratory focus on 3 major areas of synthetic biology of P. putida: tool development, tuning metabolism and production, and engineering new features.
With our decades of experience with P. putida, our laboratory has introduced novel features in P. putida and continues to create new solutions for the application of science into industrial and environmental fields. We are not only developing tools for biotechnology research but are also building new strains. For example our TNT sensing P. putida strain: This strain (which is under development) is designed such that it can report the environmental contaminants that come from explosives that are used for road constructions and so on. Another example is the P. putida strain that has unique features to be used in industrial applications as a result of modifications we have made to its chromosome. This strain is regarded as a microbial cell factory, meaning that it is an ideal chassis for producing a chemical of interest with a high yield in industry.
Apart from assigning new tasks to bacteria, in our laboratory we also work on increasing the efficiency of P. putida. This includes, for instance, tuning the metabolic activity of P. putida to use new carbon sources (from which bacteria produce energy) that it does not naturally prefer. This gives us the flexibility to use the same bacteria with different sugars. It is especially important when we consider that in industrial applications there can be different needs on the type of carbon source being used.
As a laboratory at the intersection of fundamental and application-based investigations, our focus is to discover the best technologies that have a major impact on synthetic biology and its industrial applications to promote European science and benefit society as a whole.
4. Eidgenoessische Technische Hochschule Zuerich
 Eidgenoessische Technische Hochschule Zuerich
Eidgenoessische Technische Hochschule Zuerich
Basel, Switzerland
ETH Zurich is one of the premier European research universities with an outstanding research record (including 21 Nobel laureates connected to it) and a current complement of 400 professors. Key strengths of the university are in life sciences, chemistry, engineering, and computer science. The Department of Biosystems Science and Engineering (D-BSSE) of ETH Zurich in Basel was founded in 2009 to lead a unique interdisciplinary research initiative on the interface of life sciences and engineering with two strategic thrusts, Synthetic Biology and Systems Biology. Its core conceptual thrust is to explore and engineer molecular networks in biology and biotechnology on a systems level concomitantly with the required overarching conceptual and technological developments.
 Prof. Dr. Sven Panke
Prof. Dr. Sven Panke
Main Contact
Sven Panke is a Professor of Bioprocess Engineering at the ETH Zurich. After his PhD, also at ETHZ, he worked for two years in the biocatalysis group of the Dutch chemical company DSM (Geleen, The Netherlands). He returned to ETH in 2001 as an Assistant Professor, received tenure in 2007, and then moved to the newly founded ETHZ Department of Biosystems Science and Engineering in Basel. His main research topics include integrated reaction-separation systems, high-throughput screening, and synthetic biology. His work was awarded with the ETH Medal and the DSM Research Award.
The Bioprocess Laboratory (BPL, Prof. Sven Panke) is part of the Department of Biosystems Science and Engineering (D-BSSE) of ETH Zurich. The D-BSSE is located in the tri-border area of Basel, the European “capital” of the pharma- and life science industry.
We develop novel processes for the production of biopharmaceuticals and high-value chemicals via the utilization of microbial catalysts. For engineering of improved or novel biocatalysts, molecular and synthetic biology methods such as genome editing and directed evolution are employed. Each method easily generate millions to billions of different catalyst variants (on the DNA level), which leads to another main part of the BPL’s know-how: high-throughput (HT) screening.
HT screening methods have to be co-developed during actual biocatalyst engineering projects, in order to match the possibilities of library building offered by synthetic biology. For instance, if one created a million of catalyst variants and needed “only” five minutes to assess a single variant, one would spend close to 500 weeks (almost a decade!) assessing all of them. Fortunately, basic HT screening methods already exist for more than a decade and allow the assessment of thousands of variants per second, usually based on measuring a fluorescence signal correlated with the performance of the biocatalyst (fluorescence assisted cell sorting, FACS). As a result, the time required to assess millions of catalyst variants is reduced to minutes or hours. Along these lines, the BPL investigates novel HT screening methods by developing biosensors that couple analytically inconspicuous products of interest to fluorescent signals and also investigates alternative ways of culturing microbial catalysts. This led to the development of so-called nanoliter reactors (small alginate beads harboring microbial cells), which can also be sorted into fractions based on the fluorescence of the microbial cells (Meyer et al., Nat Chem, 2015, 7: 673-78).
For the EmPowerPutida project, the BPL develops and implements metabolic engineering methods for the uncoupling of microbial cell growth and product formation, which would allow making better use of provided (and costly) nutrients. The target is to have bacterial cultures reach a sufficiently high cell density, but then should switch to a “production only” state without experiencing the common drop in productivity that typically is a side effect of no- or slow-growth regimes. To give a more graphical example: A little fruit tree would be allowed to grow for a few years to reach a certain size (biomass) that allows it to bear a lot of fruits (product), and would be kept at this size and then channel nutrients and energy into making fruits at a rate and volume equivalent to a large tree, while remaining at a handy size for gardening.
As of now, it remains unclear which functions in a cell have to modulated how to in order to reduce allocating nutrients to growth and instead channel them to product formation while keeping formation rates high. Therefore, in order to find the best options, a screening approach is applied, which needs careful engineering. Here, we exploit two strategies: Together with partner CSIC we work on methods that interrupt existing protein networks by selective proteolysis, and together with partner ITQB we develop mRNA-based methods to modulate protein translation. The identification of suitable network targets is performed in collaboration with our partners at the University of Wageningen.
The entire approach is implemented for the formation of small molecule alcohols, which can be used as biofuels and building blocks for the chemical industry. In order to facilitate the detection of such products in a HT fashion as described above, we develop biological sensing circuits that couple product titers to the formation of a fluorescent output signal. This read out is then applied to biocatalysts in nanoliter reactors under different nutrient conditions, while simultaneously and combinatorically modulating increasingly large fractions of the metabolic network.
5. Instituto de Tecnologia Química e Biológia - Universidade Nova de Lisboa
 Instituto de Tecnologia Química e Biológia - Universidade Nova de Lisboa
Instituto de Tecnologia Química e Biológia - Universidade Nova de Lisboa
Oeira, Portugal
The Instituto de Tecnologia Química e Biológica (ITQB) is a research and advanced training institute of the Universidade Nova de Lisboa. Its mission is to develop high-quality research in chemistry and the life sciences, considering all levels of complexity and their potential applications, so as to contribute to the understanding of life’s mechanisms. Its highly multidisciplinary nature makes ITQB a leading centre for advanced training of researchers in Portugal. ITQB hosts a number of independent laboratories grouped in five research divisions and consisting of a scientific staff of about 400 researchers with different scientific interests and backgrounds. Researchers at ITQB benefit from outstanding facilities, equipment, and support services, some of which are unique in Portugal.
 Prof. Dr. Cecília Arraiano
Prof. Dr. Cecília Arraiano
Main Contact
The main research area of Prof. Cecília Arraiano’s laboratory has been to elucidate determinant factors for the control of gene expression. Biological processes cannot be fully understood without a deep understanding of RNA metabolism. The main focus of the Gene Expression Lab is the study of RNA degradation mechanisms and the characterisation of enzymes that mediate RNA decay in microorganisms. Namely, ribonucleases involved in maturation, degradation and quality control of mRNAs and functional non-coding sRNAs. These studies have been applied to areas of Biotechnological interests and Health. Other research interests include bacterial stress response, cell growth and survival.
Small RNAs as a toolbox to modulate microbial metabolism
Biological processes cannot be fully understood without a detailed knowledge of RNA metabolism. All living organisms contain in their genetic code a set of rules that define how DNA is translated into proteins. In the “renaissance” of microbiology it is now clear that RNA is much more than just an intermediate, which helps to translate DNA into proteins. RNA is also a prime molecule in the control of gene expression, since its stability and vulnerability impact on protein expression yields. Furthermore, a plethora of non-coding small RNAs (sRNAs) orchestrates cell metabolism for a faster adaptation to changes in the environment.
At ITQB NOVA (Lisbon, Portugal), in the Control of Gene Expression Laboratory headed by Prof. Dr. Cecília Arraiano, we are focused on the study of RNA molecules, RNA degradation mechanisms and the enzymes that degrade RNA molecules (known as ribonucleases). Our laboratory also studies sRNAs, effectors of bacterial adaptation to changes in the outside world.
Our research has many potential applications in industrial biotechnology, since the stability of prokaryotic RNAs is a main obstacle to the profitable production of proteins of interest from industrial microorganisms. The development of programmable genetic regulators, will be a main tool in synthetic biology.
In the EmPowerPutida project, we are using synthetic biology to reprogram Pseudomonas putida to fulfill industrial production needs. We have described a pool of sRNAs, which are switched on by industrial bioreactors. Furthermore, we have constructed and are in the process of characterizing novel strains with different ribonucleases’ expression levels. Synthetic sRNAs are being built to be implemented as a tool for genetic and metabolic engineering, and their application will allow production of valuable compounds like fuels, rubber and plastic components, as well as molecules with agriculture applications. For that, we are taking advantage of some of the sRNAs features such as, modularity and composability, relative ease of design, fast dynamics and flexible tunability.
Our team has a vast expertise in in vivo and in vitro RNA methodologies, a combined knowledge that gives us advantage to establish experience-based rules for targeting essential genes involved in product formation. This will ultimately contribute to novel sRNA-based regulation devices leading to finely controlled synthetic circuits. Our findings in frame of EmPowerPutida project are of biotechnological significance and will certainly improve industrial production.

Lisbon EmPowerPutida team. From left to right: Patrícia Apura MSc, Margarida Saramago PhD, Susana Domingues PhD, Prof. Dr. Cecília M. Arraiano, Sandra Viegas PhD and Pseudomonas putida KT2440.
6. Ingenza Limited
 Ingenza Ltd.
Ingenza Ltd.
Edinburgh, United Kingdom
Ingenza: An Industrial Biotechnology Company with a broad customer base across the chemicals, pharmaceuticals, food, feed and fuel industries. We apply synthetic biology to the manufacture of industrial products including enhanced biofuels, sustainable manufacturing of chemicals and the production of protein therapeutics. Our scientific and commercial activities are led by a management team with over 25 years’ experience in applied bioscience and the development and commercialisation of biobased products.
In addition to engaging in strategic partnerships to tailor our bioprocess services for clients, we also license our proprietary bioprocess technologies.
 Dr. Ian Fotheringham
Dr. Ian Fotheringham
Main Contact
Ian Fotheringham received a Ph.D. in Molecular Biology from the University of Glasgow, UK in 1986. He joined the NutraSweet division of Monsanto in Chicago, USA, constructing microbes to produce the Aspartame® sweetener. From 1993 he continued developing large scale bioprocesses with NSC Technologies and Great Lakes Fine Chemicals. In 2003 he co-founded Ingenza, an Edinburgh, UK, based Industrial Biotechnology SME with a unique range of proprietary enabling technologies. Now a leader in microbial strain improvement, synthetic biology, fermentation and bioprocess development, Ingenza works with industrial partners worldwide to commercialise state of the art biomanufacturing processes. Ian has published 35 papers and articles and holds 8 current patents.
7. LifeGlimmer GmbH
 LifeGlimmer GmbH
LifeGlimmer GmbH
Berlin, Germany
LifeGlimmer GmbH is an international science-driven company that caters advanced solutions for the full research pipeline in bioinformatics, data integration, and systems biology. From designing workflows, via sophisticated analytics and dynamic simulations, to new knowledge discovery and novel inroads to applications in both fundamental and applied biological and biomedical practice, LifeGlimmer GmbH offers tailored scientific solutions to deal with complexity in the biosciences. In addition LifeGlimmer GmbH seeks to advance scientific projects with services and consulting in project management, research strategy development, experimental design and specialised training.
Mechthild Schulte M.A., has outstanding experience in business administration and public relations in international environments. As science manager in the public sector she gained profound expertise in the development and controlling of EC and nationally funded research projects. She joint LifeGlimmer Gmbh as chief administrator and involves in the cooperation of scientific and economic bodies.
M.Sc. François Bergey graduated in statistical mathematics. He joint LifeGlimmer GmbH in 2015. He applies his expertise in time-series RNA seq data, data mining of clinical data, text mining in medical health records and predictive modeling for experimental biology and health related science. He is also experienced in Bioinformatics, performing pathway analysis, graph search and data management of large dataset.
8. Abengoa Research SL
 Abengoa Research SL
Abengoa Research SL
Seville, Spain
Abengoa is a world leader in the development of innovative technology solutions for a sustainable development. Abengoa’s R&D is centralised in Abengoa Research which is a company belonging to Abengoa. The core objective is to generate knowledge and promoting its applications in the field of energy and sustainable development. Abengoa organises its research, development and innovation into the following areas: materials and nanotechnology, fluid mechanics, mechanics of solids and structures, thermal engineering, process engineering, biotechnology and biomass, and electrical networks.
Abengoa Research is developing new approaches to build synthetic pathways for biosynthesis of fuels and added-value chemicals.
9. BASF SE
 BASF SE
BASF SE
Ludwigshafen, Germany
At BASF, we create chemistry for a sustainable future. We combine economic success with environmental protection and social responsibility. The approximately 114,000 employees in the BASF Group work on contributing to the success of our customers in nearly all sectors and almost every country in the world. Our portfolio is organized into five segments: Chemicals, Performance Products, Functional Materials & Solutions, Agricultural Solutions and Oil & Gas. BASF generated sales of about €58 billion in 2016. BASF shares are traded on the stock exchanges in Frankfurt (BAS), London (BFA) and Zurich (BAS).
 Dr. Andrea Herold
Dr. Andrea Herold
Main Contact
Dr. Andrea HEROLD has worked for BASF for more than 10 years. She worked in various functions in BASF’s biotechnology research and New Business Development and has a strong background in metabolic engineering. Currently, she is group leader of the Microbiology group, responsible for the development of microbial production processes of biopolymers, biological crop protection products and secondary metabolites.
Dr. Lutz Petzke is a research scientist in the Microbiology group at BASF. His expertise is the microbial production of secondary metabolites. He has more than 5 years of experience in engineering Actinomycetes and Pseudomonas strains with a strong focus on biosynthetic cluster expression.
With her background in industrial biotechnology she has been developing strains and processes for the fermentative production of bio-based chemicals for 4+ years. During that time she has worked with various industrially relevant microorganisms on different target molecules. Her current focus is on secondary metabolites production.
10. Lucite International
Lucite International
Billingham, United Kingdom
Lucite International is the world's leading supplier of Methyl Methacrylate (MMA), and a global leader in the design, development and manufacture of acrylic based products. It is the only organization to manufacture MMA with production, R&D, sales and marketing facilities in all three major geo-economic regions; the Americas, Europe and Asia. The Company employs 2000 people to develop, manufacture and sell acrylic based products to customers in more than 100 countries. Lucite International has 14 manufacturing sites and 34 sales offices around the globe. Lucite International is and brings decades of expertise with the petrochemical base manufacture of the target products and great depth in the chemistry to produce the target molecules from alternate, bio-derived starting materials. Lucite began the manufacture of MMA as early as the 1930’s when the material was used in the manufacture of spitfire cockpit canopy's. The company developed as part of the ICI group of companies and achieved its place as the number one producer as a result of the acquisition of the Dupont acrylics business in the early 1990’s. More recent growth was achieved under the guidance of the Charterhouse group of companies as ICI divested many of its bulk chemicals operations in the late 1990’s, and after acquisition in 2008, the company is now a wholly owned subsidiary of Mitsubishi Chemical Corporation. Despite these changes in ownership the acrylics business has continued to grow at a rate in excess of average GDP due mainly to the excellent properties of PMMA and the success of Lucite and its customers in finding new applications for this material. Lucite is fully committed to producing MMA via a bio-based route. We live in a rapidly changing world where fossil fuels are recognised to be finite resource and where the future of chemicals production will be increasingly dominated by plant-based feedstocks. Methacrylates are a strong candidate to be part of this future bio-based chemical industry, especially from the sustainability perspective, since they have good weatherability and can be easily recycled to generate the starting monomer. Although direct bio-based routes to methacrylate monomers do not exist at present, the area of producing methacrylates via biotechnology is becoming increasingly competitive.
 Dr. Bastian Blombach
Dr. Bastian Blombach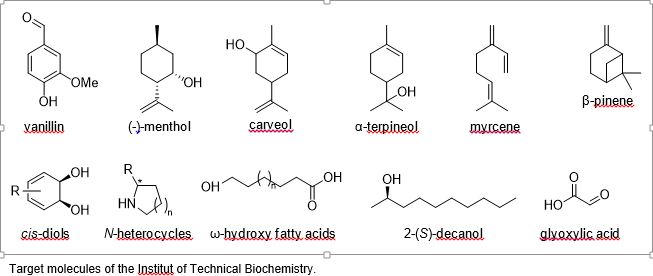
 Mechthild Schulte M.A.
Mechthild Schulte M.A.  M.Sc. François Bergey
M.Sc. François Bergey


